 Cross-posted from Nothing in Biology Makes Sense!
Cross-posted from Nothing in Biology Makes Sense!
It is a truth universally acknowledged in evolutionary biology, that one species interacting with another species, must be having some effect on that other species’ evolution.
Actually, that’s not really true. Biologists generally agree that predators, prey, parasites, and competitors can exert natural selection on the other species they encounter, but we’re still not sure how much those interactions matter over millions of years of evolutionary history.
On the one hand, groups of species that are engaged in tight coevolutionary relationships are also very diverse, which could mean that coevolution causes diversity. But it could be that the other way around: diversity could create coevolutionary specificity, if larger groups of closely-related species are forced into narower interactions to avoid competing with each other.
Part of the problem is that it’s hard to study a species evolving over time without interacting with any other species—how can we identify the effect of coevolution if we can’t see what happens in its absence? If only we could force some critters to evolve with and without other critters, and compare the results after many generations …
Oh, wait. That is totally possible. And the results have just been published.
A team of evolutionary microbiologists has performed exactly the experiment I outlined above. The study’s lead author is Diane Lawrence, a Ph.D. student in the lab of Timothy Barraclough, who is listed as senior author.
For the experiment, the team isolated five bacterial species, of very different lineages, from pools of water at the bases of beech trees—ephemeral pockets of habitat for all sorts of microbes that break down woody debris, dead leaves, and other detritus. They cultured the bacteria on tea made from beech leaves, in vials containing either a single species, or all five species, and let them evolve for eight weeks—several dozens of bacterial generations. In a particularly clever twist on standard experimental evolution methods, they also used nuclear magnetic resonance (NMR) to identify the carbon compounds in sterilized tea that had been “used up” by the bacterial cultures, and compared the compounds in fresh beech tea to determine what the bacteria were eating.
 The base of a beech tree. Photo by -nanio-.
The base of a beech tree. Photo by -nanio-.And, maybe not surprisingly, the bacterial species’ evolution with company turned out to be quite a bit from their evolution alone. Left alone, most of the species evolved a faster growth rate. This is a common result in experimental evolution, because the process of transferring evolving bacteria to fresh growth medium—”serial transfers” that were performed fifteen times over the course of the experimetn—can create natural selection that favors fast-growing mutants. But, grown all together in the same tube, species that had evolved faster growth rates in the solo experiment evolved slower growth instead.
To find out what had evolved in the multi-species tubes, the team tested the growth of the bacterial species on beech tea that had been used to grow one of the other species, then sterilized. The original, ancestral strains of bacteria generally had negative effects on each others’ growth—they lived on similar compounds in the beech tea, and so their used tea wasn’t very nourishing for the other species. The same thing occurred with the strains that had evolved alone, only stronger, which makes sense in light of the increased growth rates, which would’ve depleted the growth medium faster.
But the interactions among the strains of the different bacterial species that had evolved together was strikingly different. Many of them actually made the tea more nutritious for other species in the evolved community. That is, some of the bacteria had evolved the capacity to eat the waste products of another species that was evolving with them. Using the NMR method to track changes in the presence of different carbon compounds in the tea before and after use provided confirmation that the co-evolved species were using, and producing, complementary sets of resources.
In short, the evolving community didn’t simply become more diverse—it evolved new kinds of mutually beneficial relationships between species that began as competitors.
That evolutionary shift toward mutual benefit had a significant impact on the bacterial community as a whole, too. Lawrence et al. assembled new communities of bacteria extracted from the end-point of the group evolution experiment, and compared their carbon dioxide production, a proxy for overall metabolic activity, to that of a community assembled from bacteria extracted from the end point of the solo-evolution experiments. The community of co-evolved bacteria produced significantly more carbon dioxide, suggesting they were collectively able to make more use out of the growth medium.
So that’s a pretty nifty set of results, I have to say. But I’m also left wondering what it tells us more generally. In both Lawrence et al.‘s paper, and in accompanying commentary by Martin Tucotte, Michael Corrin, and Marc Johnson, there’s a fair bit of emphasis on the unpredictability of the result. Lawrence et al. write, in their Discussion section,
The way in which species adapted to new conditions in the laboratory when in monoculture—the setting assumed for many evolutionary theories and experiments—provided little information on the outcome of evolution in the diverse community.
And, as Corrin et al. note,
These results imply that predictions constructed from single-species experiments might be of limited use given that most species interact with many others in nature.
So … evolution went differently under different conditions? That isn’t exactly a shocking revelation. The fact that this is one of the study’s major conclusions is a symptom of how little experimental work has actually tested the effects of multiple species on evolution. One experiment I’ve discussed here previously, focused on the joint effects of predators and competitors on microbes that live in pitcher plant pitfalls, similarly emphasized the fact that it wasn’t possible to predict the evolutionary effects of predators and competitors together based solely on their individual effects. Work in this line of inquiry is hanging at the point of establishing that complex conditions lead to complex results.
What I’d really like to know—and I think all the authors of both the paper and the commentary would agree with me on this—is how we can begin to make general predictions about community evolution beyond, “it depends what we put in at the start.” It may be that we’ll need a lot more studies like this current one before we can start to identify common processes, and more interesting trends.◼
References
Turcotte, M., Corrin, M., & Johnson, M. (2012). Adaptive evolution in ecological communities. PLoS Biology, 10 (5) DOI: 10.1371/journal.pbio.1001332
Lawrence, D., Fiegna, F., Behrends, V., Bundy, J., Phillimore, A., Bell, T., & Barraclough, T. (2012). Species interactions alter evolutionary responses to a novel environment. PLoS Biology, 10 (5) DOI: 10.1371/journal.pbio.1001330
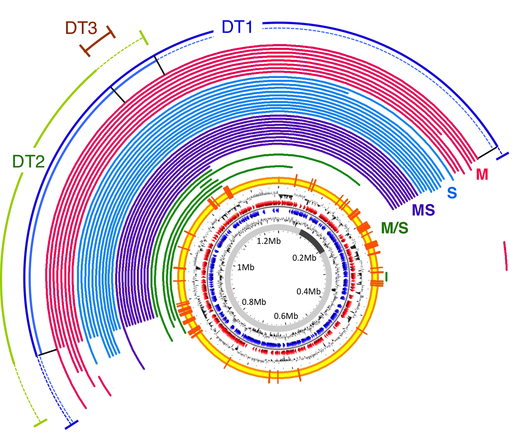 Image via Nothing in Biology Makes Sense.
Image via Nothing in Biology Makes Sense.




